Introduction: Cerebral palsy (CP) is the most common physical disability in childhood characterized by irreversible, non-progressive central motor dysfunction. Genetic factors, and environmental insults such as prematurity and hypoxia-ischemia, can contribute to CP. However, significant gaps in our knowledge of the genetic basis of CP impede advances in preventive, diagnostic, and therapeutic measures. We recently performed whole-exome sequencing (WES) in 250 CP trios and observed that at least 14% of CP cases could be accounted for by damaging genomic variants, and many of these CP risk genes control the wiring process of brain circuits in early development (Jin et al., Nat. Genet. 2020). Although our results revealed a strong genetic contribution to CP, the causes for most patients remain largely unknown.
Methods: Our objectives are to provide mechanistic insight into newly identified genetic causes and make genotype-phenotype correlations by applying an integrative, multi-dimensional omics approach to a large, well-phenotyped CP cohort. We thus hypothesize: (1) a proportion of the unexplained CP cases could be explained by de novo single nucleotide variants (SNVs), de novo copy number variations or structural variations that could only be detected by whole-genome sequencing (WGS), and (2) high-throughput functional assays can facilitate interpretation of these genetic variants.
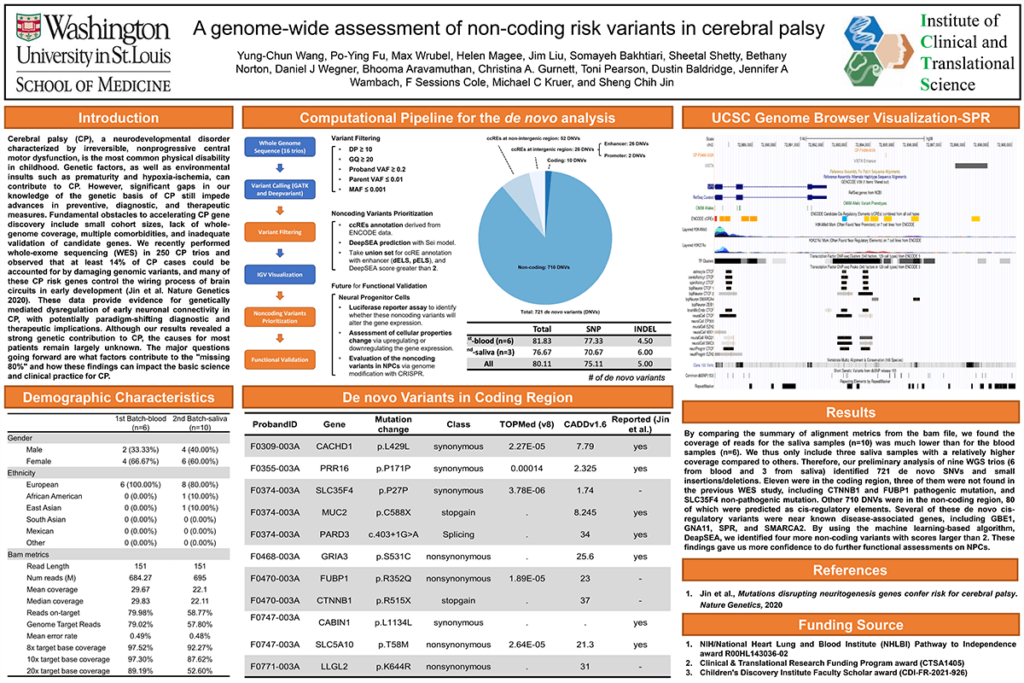
Results: WGS has been completed on ~50 cerebral palsy trios in whom WES failed to identify pathogenic variants from the Undiagnosed Diseases Network and Cerebral Palsy Research Network. Our preliminary analysis of six WGS trios identified 492 de novo SNVs and small insertions/deletions. Eight were in the coding region, one of which being a CTNNB1 pathogenic mutation that was not found in the previous WES study. Other 484 DNVs were in the non-coding region, 56 of which were predicted as cis-regulatory elements. Several of these de novo cis-regulatory variants were near known disease-associated genes, including GBE1, GNA11, SPR, and SMARCA2.
Impact: Our finding suggested potentially disruptive cis-regulatory elements could explain a fraction of WES-negative CP patients.
Organization – Washington University in St. Louis
Ali L, Patel P, Vakaki M, Kremitzki C, Waligorski J, Chadrasekaran V, Kelley K, Bachman G, Wang YC, Fu PY, Wrubel M, Magee H, Liu J, Bakhtiari S, Shetty S, Norton B, Wegner DJ, Aravamuthan B, Gurnett CA, Pearson T, Baldridge D, Wambach JA, Cole FC, Kruer MC, Jin SC